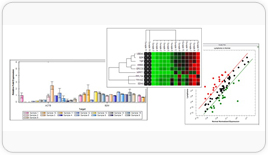

Matrix normalized worksheet reports normalized signal (against housekeeping genes).
#BIO RAD CFX MANAGER GENE STUDY SOFTWARE#
The CFX manager software automatically performs all deltadeltaCt based fold-change calculations from the uploaded raw threshold cycle data.įold Change worksheet reports test/control (CS-MSCs/BM-MSCs) ratios.Ĭomparative analysis of human bone marrow-derived MSCs and corneal stromal cells Target gene signals normalized to housekeeping genes 2^-deltaCt, where deltaCt = (Ct_Target − Ct_HKG)] The normalization and all the data analysis were performed according to the manufacturers instructions using the CFX Manager software Gene Study ( )įor the normalization it uses the average of three housekeeping genes: TBP, HPRT1 and GAPDH. Quantitative real-time PCR were performed (CFX96 Real-Time System – C1000 Thermal Cycler, Bio-Rad) with 40 cycles at 95☌ for 5 seconds and 60☌ for 30 seconds. From the purified RNA, 10ng was used per sample to synthesize cDNA using the iScript Advanced cDNA synthesis kit (Bio-Rad Laboratories, Temse, Belgium) following the protocol supplied by the manufacturer. Create a gene study to compare gene expression data from one or more real-time PCR experiments using an inter-run calibrator to normalize between the. QRT-PCR assays were prepared in triplicate in 96-well PrimePCR plates for human MSCs (Mesenchymal stem cells, SAB Target List, Bio-Rad Laboratories, Temse, Belgium). Total RNA was extracted with RNeasy Mini Kit followed by DNase I treatment. All cells were used in the experiments before the 6th passage. When 80% to 90% confluence was reached, cells were passaged using TrypLE (Life Technologies, Gent, Belgium) for 5 minutes. BM-MSCs from a single donor were expanded in DMEM + 10% FBS. Candidate Gene-Based Association Mapping of ZmPLC Family Members in Maize. Data were analyzed using Bio-Rad CFX Manager software. For all qRT-PCR analyses, triplicate biological samples were collected. GEO help: Mouse over screen elements for information.Ĭell type: corneal stromal mesenchymal stem cellsĬS-MSCs from six donors were seeded after isolation in both Dulbecco’s Modified Eagle Medium (DMEM) supplemented with 10% Fetal Bovine Serum (FBS), as the standard culture condition, and in DMEM supplemented with 10% Human Platelet Lysate (HPL), following our optimized xeno-free culture protocol reported previously (Matthyssen et al. The reaction was performed on Bio-Rad CFX Connect TM using SYBR-Green to detect gene expression levels.
